|
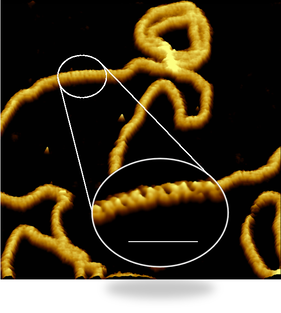
In a study published in PNAS, we have reported some of the sharpest images ever of entire living bacteria, revealing the complex architecture of the protective layer that surrounds many bacteria and makes them harder to be killed by antibiotics. Our study, carried out in collaboration with scientists at National Physical Laboratory, King’s College London, University of Oxford and Princeton University, reveals that bacteria with protective outer layers – called Gram-negative bacteria – may have stronger and weaker spots on their surface.
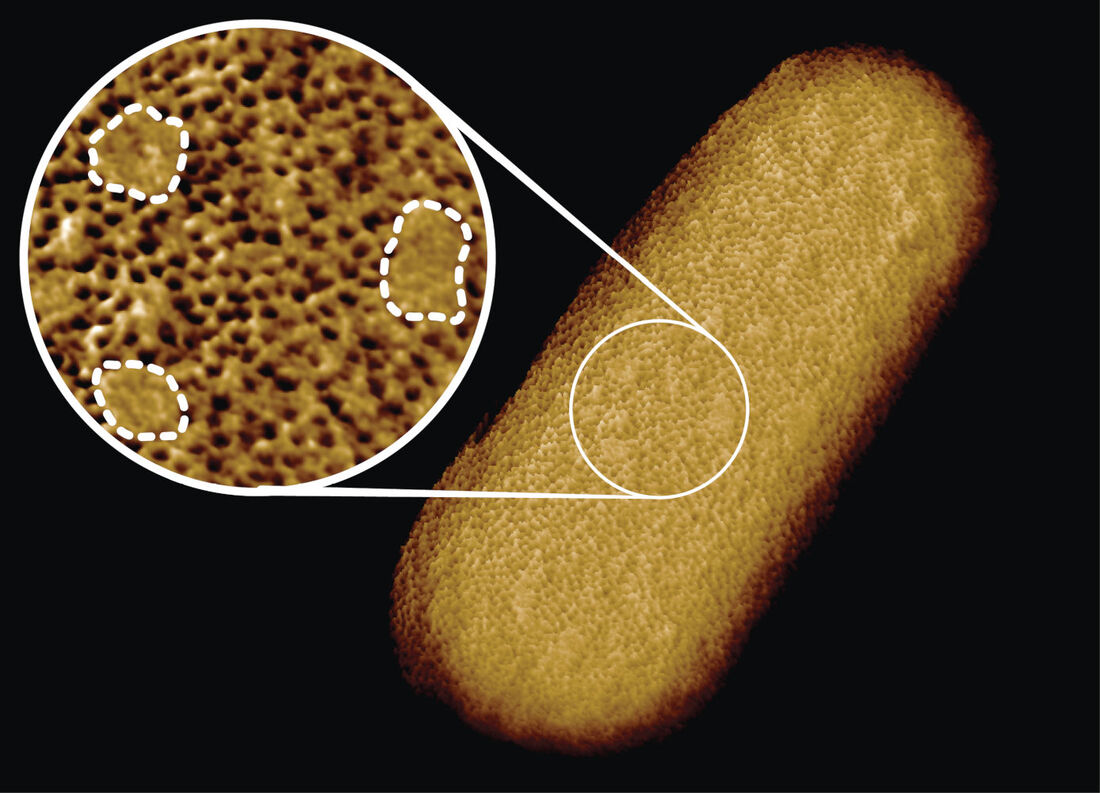
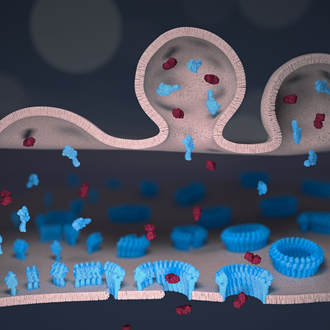
We found that the protective outer membrane of the bacteria contains dense networks of protein building blocks alternated by patches that do not appear to contain proteins. Instead, these patches are enriched in molecules with sugary chains (glycolipids) that keep the outer membrane tight. This is an important finding because the tough outer membrane of Gram-negative bacteria prevents certain drugs and antibiotics from penetrating the cell: this outer membrane is part of the reason why antimicrobial resistance of such bacteria (including A. baumannii, P. aeruginosa, and enterobacteriaceae such as Salmonella and E. coli) is now considered a greater threat than that of Gram-positive bacteria such as resistant S. aureus (well known as MRSA). “The outer membrane is a formidable barrier against antibiotics and is an important factor in making infectious bacteria resistant to medical treatment. However, it remains relatively unclear how this barrier is put together, which is why we chose to study it in such detail,” explained corresponding author Professor Bart Hoogenboom (London Centre for Nanotechnology at UCL and UCL Physics & Astronomy). “By studying live bacteria from the molecular to cellular scale, we can see how membrane proteins form a network that spans the entire surface of the bacteria, leaving small gaps for patches that contain no protein. This suggests that the barrier may not be equally hard to breach or stretch all over the bacterium, but may have stronger and weaker spots that can also be targeted by antibiotics.” To better understand this architecture, the scientists ran a tiny needle over living Escherichia coli (E. coli) bacteria, thus “feeling” their overall shape. Since the tip of the needle is only a few nanometres wide, this made it possible to visualise molecular structures at the bacterial surface. The resulting images show that the whole outer membrane of the bacteria is crammed with microscopic holes formed by proteins that allow the entry of nutrients while preventing the entry of toxins. Although the outer membrane was known to contain many proteins, this crowded and immobile nature had been unexpected. Surprisingly, the images also revealed many patches that did not appear to contain proteins. These patches contain a glycolipid normally found on the surface of Gram-negative bacteria. In addition, a different type of pimple-like patch formed when parts of the membrane were flipped inside out due to mutations. In this case, the appearance of these defects correlated with enhanced sensitivity to bacitracin, an antibiotic usually only effective against Gram-positive, but not against Gram-negative bacteria. As explained by Georgina Benn, who did the microscopy on the bacteria in Professor Hoogenboom’s lab at UCL: “The textbook picture of the bacterial outer membrane shows proteins distributed over the membrane in a disordered manner, well-mixed with other building blocks of the membrane. Our images demonstrate that that is not the case, but that lipid patches are segregated from protein-rich networks just like oil separating from water, in some cases forming chinks in the armour of the bacteria. This new way of looking at the outer membrane means that we can now start exploring if and how such order matters for membrane function, integrity and resistance to antibiotics.” We speculate that the findings may help explain ways by which bacteria can maintain a tightly packed, protective barrier while still allowing rapid growth: the common bacterium E. coli doubles in size and then divides in 20 minutes under favourable conditions. Specifically, the glycolipid patches may allow for more stretch of the membrane than the protein networks, making it easier for the membrane to adapt to the growing size of the bacterium. The work was kindly funded by the UKRI, the National Institutes of Health, the European Research Council, and the UK Department for Business, Energy and Industrial Strategy. 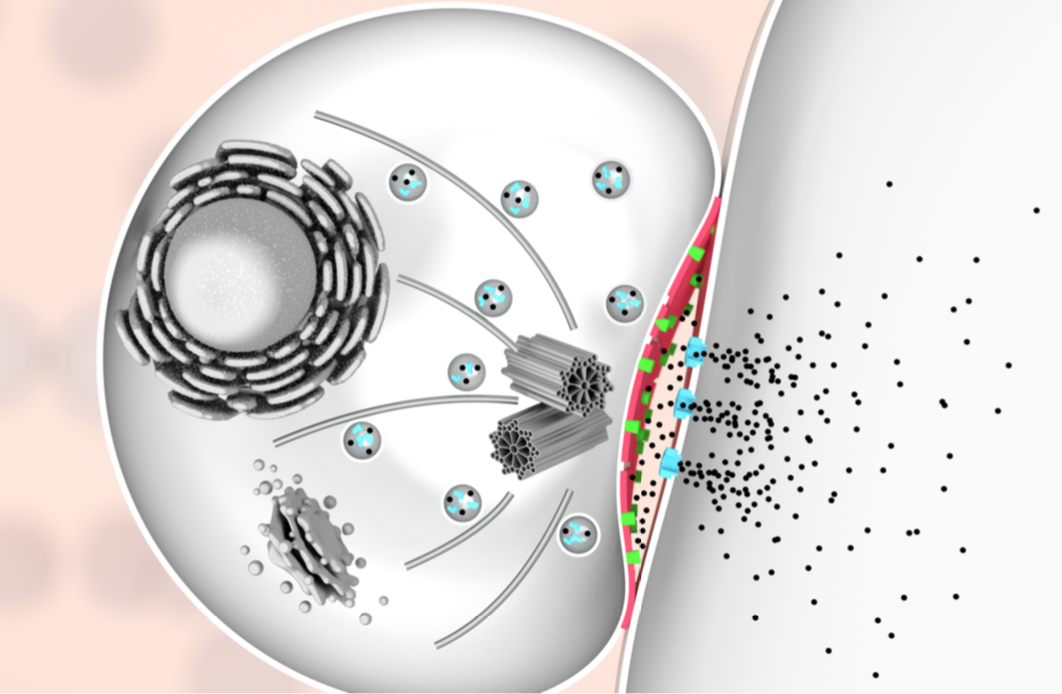 To clear our body from cancerous and virus-infected cells, our immune system relies on a type of white blood cells called killer lymphocytes. Once they have identified their targets, these white blood cells release a toxic potion of proteins. In a study published in Nature Communications, we have found what prevents the white blood cells from being killed by their own actions. Our findings could help to explain why some tumours are more resistant than others to recently developed cancer immunotherapies. Cytotoxic lymphocytes rid the body of disease by punching holes in rogue cells and by injecting poisonous enzymes inside. Remarkably, they can do this many times in a row, without harming themselves. Our study, carried out in tight collaboration with Ilia Voskoboinik’s lab at the Peter MacCallum Cancer Centre in Melbourne, discovers the protective role of molecular order and electric charge in the cell envelope of the white blood cells. We made this discovery by studying perforin, which is the protein responsible for the hole-punching. We found that perforin’s attachment to the cell surface strongly depends on the order and packing of the molecules – so-called lipids – in the membrane that surrounds and protects the white blood cells. More order and tighter packing of the lipid molecules led to less perforin binding, and when we artificially disrupted the order of this lipid in the white blood cells, the cells became more sensitive to perforin. However, we also found that when the white blood cells were exposed to so much perforin that some of it stuck to their surface, the bound perforin still failed to kill the white blood cells, indicating that there must be another layer of protection. This turned out to be the negative charge of some lipid molecules sent to the cell surface, which bound the remaining perforin and blocked it from damaging the cell. According to Adrian Hodel, joint first author and PhD student in our lab: “We have long known that local lipid order can change how cells communicate which each other, but it was rather surprising that the precise physical membrane properties can also provide such an important layer of protection against molecular hole-punchers.” In Melbourne, joint first author Jesse Rudd-Schmidt, who focused on the characterisation of the white blood cells in Ilia Voskoboinik’s laboratory, said: “What we have found helps to explain how our immune system can be so effective in killing rogue cells. We are now also keen to investigate if cancer cells may use similar protection to avoid being killed by immune cells, which would then explain some of the large variability in patient response to cancer immunotherapies.” The work was kindly funded by the Australian National Health and Medical Research Council and by the UK Biotechnology and Biological Sciences Research Council and Engineering and Physical Sciences Research Council, and by the Sackler Foundation. One of the early breakthroughs for nanotechnology was its success in visualising individual atoms and molecules. Nowadays, we can not only visualise molecules, but also capture videos of how they move. Yet at the nanoscale, how do you know if such motion is related to molecular dynamics, and not simply due to measurement noise? In a study published in the journal ACS Nano, we have developed methods to answer this question.
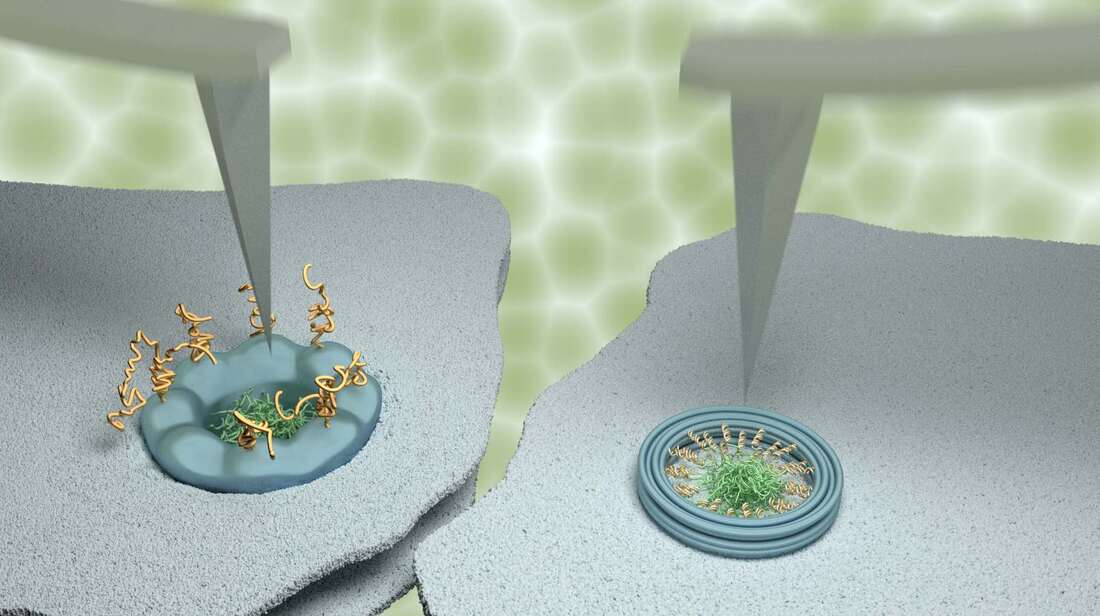
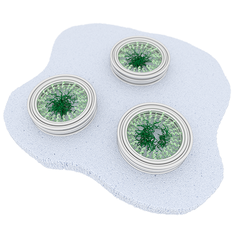
Our study made extensive use of a technique named atomic force microscopy. This type of microscopy uses an ultrafine needle to feel rather than see molecules on a surface, similar to a blind person reading Braille. The needle repeatedly scans the surface to produce an image that refreshes fast enough to track how molecules or groups of molecules change with time. To demonstrate the power of our methods, we focussed on tiny channels in the protective membrane that surrounds the nucleus in our cells. For the appropriate use of genetic material in the nucleus, these ‘nanopores’ selectively allow the entrance and exit of certain molecules, while blocking others to protect genetic material and normal functions of the cell. The results were compared with data obtained on artificial nanopores – built by collaborators at Yale University – that replicate several properties of the natural nanopores in the nuclei of our own cells. Whereas the artificial nanopores showed unambiguous signatures of molecular motion, this was much less clear for the natural pores. In agreement with results previously reported by some other labs, some (sub)nanometre fluctuations were detected; yet upon closer inspection, these fluctuations turned out to be indistinguishable from fluctuations measured away from the nanopores, and were therefore attributed to measurement noise. We explain the difference between the natural and artificial nanopores by the presence of other proteins that are involved in the transport and that in the natural nanopores act as a glue preventing or restricting the motion of other molecules. This work was kindly supported by the Biotechnology and Biological Sciences Research Council (BBSRC) and by the Engineering and Physical Sciences Research Council (EPSRC). Joint first authors and PhD students George Stanley and Bernice Akpinar are part of, respectively, the BBSRC London Interdisciplinary Doctoral Programme (LIDo) and the joint UCL/Imperial College London Centre for Doctoral Training in the Advanced Characterisation of Materials. To kill bacteria in the blood, our immune system relies on nanomachines that can open deadly holes in their targets. We have now filmed these nanomachines in action, discovering a key bottleneck in the execution of their killer function, which helps to protect our own cells.
Our research, published in Nature Communications, provides us with a better understanding of how the immune system kills bacteria and of why our own cells remain intact. This may guide the development of new therapies that harness the immune system against bacterial infections, and strategies that repurpose the immune system to act against other rogue cells in the body. In earlier research, published in EMBO Journal earlier this year, we had imaged the hallmarks of attack on live bacteria, showing that the immune system response results in ‘bullet holes’ spread across the pathogen’s cell envelopes. The holes are incredibly small with a diameter of just 10 nanometres – about 1/10,000 of the width of a human hair. To film this process in real time, we mimicked how these deadly holes are formed by the membrane attack complex (MAC) using a model bacterial surface. By tracking each step of the process, we found that shortly after each hole started to form, the process stalled, offering a reprise for the body’s own cells. “It appears as if these nanomachines wait a moment, allowing their potential victim to intervene, in case it is one of the body’s own cells instead of an invading bug, before they deal the killer blow,” explained Dr Edward Parsons, first author of this study and postdoc in our lab. The process pauses as 18 copies of the same protein are needed to complete a hole. Initially, there’s only one copy which inserts into the bacterial surface, after which the other copies of the protein slot into place much more rapidly. “It is the insertion of the first protein of the membrane attack complex which causes the bottleneck in the killing process. Curiously, it coincides with the point where hole formation is prevented on our own healthy cells, thus leaving them undamaged,” said Professor Bart Hoogenboom. To film the immune system in action at nanometre resolution and at a few seconds per frame, we used atomic force microscopy. This type of microscopy uses an ultrafine needle to feel rather than see molecules on a surface, similar to a blind person reading Braille. The needle repeatedly scans the surface to produce an image that refreshes fast enough to track how immune proteins get together and cut into the bacterial surface. The research was carried out in close collaboration with scientists from Imperial College London, from the Swiss Federal Institute of Technology in Lausanne, and from the University of Leeds; and for the work on bacteria, we relied on a collaboration with the University Medical Centre Utrecht. The work was kindly funded by the Biotechnology and Biological Sciences Research Council, Medical Research Council, Engineering and Physical Sciences Research Council and the European Union. Inspired by biological filters and in tight collaboration with collaborators at Yale University, we have designed, built and characterised a nanometre-sized pore that helps us to better understand the complex way by which cells transport genetic material and viruses.
Our study, published in ACS Nano, shows how DNA can be folded using ‘DNA origami’ to form a cylinder more than 1,000 times smaller than the width of a hair. This cylinder can then be filled by the proteins that in our body selectively transport material into and out of the nuclei of our cells, resulting in a ‘nano-pore’ filter. “We don't have a good understanding of how the complex machinery in our cells filters, for example, RNA out of and – to our detriment – viruses into the cell nucleus. Our nanopore mimics the system, but has been built from the bottom-up, allowing us to add in complexity step by step, such that we can identify the minimal requirements for nuclear import and export selectivity,” explained Bernice Akpinar, co-author of the study and PhD student in our lab, who also spent three months at Yale in the context of this project. “We were inspired by the way our body selectively protects the genetic material located in each cell nucleus,” explained co-author and PhD student Luke Davis. “This is done by spaghetti-like proteins in filters that are called nuclear pore complexes. These filters are super-selective, but as the name suggests, they are rather complex.” This collaborative study was initiated at the Yale School of Medicine by senior authors Patrick Lusk and Chenxiang Lin. Lusk notes that “For a long time, biologists have struggled to grasp how molecules are blocked from entering the cell nucleus so efficiently. Now, using DNA origami, we can make and control these bio-inspired systems and have a very powerful tool to start understanding how complex biological machinery works.” The next step for the team is to investigate how such a nanopore responds to different proteins that can or cannot pass the biological filters by which it was inspired. Then it can be embedded into a device for measuring transport and the actual filtration function. Professor Bart Hoogenboom, who supervised the UCL part of this research, said: “A few years ago, we predicted that filtering proteins would be able to switch the pores between open and closed states. It has been very rewarding to see that type of behaviour in a bio-inspired device.” In the longer term, the nanopore which acts as a nano-scale filter could be used to develop bio-inspired devices for filtering and detoxification. He added: “In terms of applications, we think molecular filters may be useful in preventing background proteins from reaching detector devices and sensing technologies, reducing background noise and improving efficacy. We also think the insights we gain on the nano-scale model system will inform the design of larger scale filters for purification, detoxification in other contexts.” The work was kindly supported by the Engineering and Physical Sciences Research Council (EPSRC) and by the National Institutes of Health (NIH). PhD student Bernice Akpinar is part of the joint UCL/Imperial College London Centre for Doctoral Training in the Advanced Characterisation of Materials. Together with researchers at National Physical Laboratory researchers (NPL), we have developed and characterised a synthetic ‘virus’ that kills bacteria on first contact.
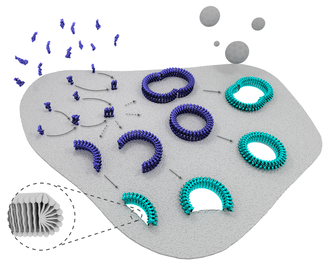
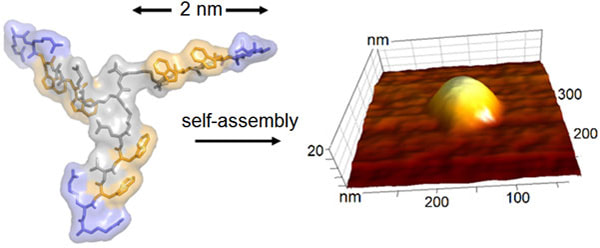
Our study, published in Nature Communications, shows how newly designed proteins can be used to build tiny hollow shells that emulate the outer structures of naturally occurring viruses. The synthetic virus ‘drones’ recognise bacterial cells before targeting and destroying their most vulnerable part – their membrane. “We used high-resolution and real-time imaging to see the impact of the synthetic viruses on bacterial model membranes and found that they are extremely destructive,” explained Hasan Alkassem, co-author and PhD student in our lab. “Seconds after landing on the surface, the synthetic viruses disassemble and form rapidly expanding holes in the membrane, causing it to leak. Experiments on intact bacteria then showed that this caused the bacteria to die.” The discovery is important as it demonstrates a possible new approach to tackling the ever-growing issue of antibiotic resistance, by opening new options for the treatment of infectious diseases. More than 700,000 people across the world die from drug resistant infections every year and antibiotic discovery has fallen well behind its historical rate, with traditional discovery methods being exhausted. “The way our synthetic virus works means that bacteria are less likely to become resistant to it. They also have an additional advantage over traditional antibiotics in that they don’t have to reach and hit a single target inside a bacterial cell to be effective,” said Alex Yon, co-author and PhD student in our lab. Furthermore, because such synthetic viruses leave human cells unaffected, but have the ability to enter them like natural viruses do, they hold promise for gene delivery and gene editing – core capabilities for gene therapy and synthetic biology – as well as for killing bacteria that hide inside human cells. Lead author, Dr Max Ryadnov, the NPL’s science leader in biometrology, explained: “This discovery adds to the growing toolbox of engineering metrology methods and materials being developed at NPL to realise the full potential of synthetic biology for industry and healthcare. This research also offers long-term and creative solutions for alternative treatments of infectious diseases that are urgently needed.” Professor Bart Hoogenboom concluded: “We were particularly pleased to find that our observations of membrane damage at nanometre length scale translated into the actual bacteria being killed. Hence we have a potential strategy to treat infectious diseases, and we also understand how it works.” The work was kindly funded by the Engineering and Physical Sciences Research Council (EPSRC), European Metrology Programme for Innovation and Research (EMPIR) and the Department for Business, Innovation and Skills (BEIS). Specialist measurements were performed at the Diamond Light Source. Our immune system can attack and destroy harmful invaders such as virus-infected and cancerous cells by punching holes in their membranes. As reported in Nature Nanotechnology, we have visualised this process in microscopic detail, in a collaboration with scientists from UCL, Birkbeck, University of London, Peter MacCallum Cancer Centre and Monash University, Australia. Our results deepen the understanding of the critical role of the protein called ‘perforin’ in a functioning immune system, bringing us one step closer to new therapies with the potential to boost or inhibit its impact when required. We used atomic force microscopy and electron microscopy to reveal precisely how a subset of white blood cells, called cytotoxic lymphocytes (or killer T cells), show remarkable efficiency in first perforating their victims and next injecting poisonous enzymes to rid the body of disease. Using a form of microscopic CCTV, we showed how perforin binds to the protective membrane that surrounds harmful cells. Professor Hoogenboom, said: “Our immune system needs to drill holes in virus-infected and cancerous cells to get rid of them, but can’t buy such drills in a DIY shop. We have now shown how it self-assembles these drills on the spot by putting multiple perforin molecules together in ring-like structures, leaving tiny holes - just tens of nanometres in diameter.” Associate Professor Ilia Voskoboinik, a lead co-author (Peter MacCallum Cancer Centre), said: “To kill virus-infected or cancerous cells, perforin must be quick and efficient. Our experiments in Melbourne show that patients who are born with impaired perforin may present with fatal failure of the immune system and also have a higher risk of developing blood cancers. “This was entirely consistent with the microscopic data obtained in London, which shows that the effectiveness of perforin is greatly hampered even if only a small number of the perforin molecules are abnormal. This new understanding brings us one step closer to targeted therapies that can strengthen the body’s perforin-producing power to ward off disease. We could also inhibit its function to prevent the rejection of organ transplants, when accepting foreign tissue or cells can be instead life-saving.” To film perforin in action, the scientists used atomic force microscopy in our lab at the London Centre for Nanotechnology. This type of microscopy uses an ultrafine needle to feel rather than see perforin on a target membrane, similar to a blind person reading Braille. The needle repeatedly scans the surface to produce an image that refreshes fast enough to track how perforin molecules get together and cut holes in the membrane. Initially, perforin appeared as a blur on these images. However, once a few perforin molecules together inserted into the membrane, they could be more clearly identified and shown to recruit more perforin to thus growing transmembrane pores. By also recording static snapshots at higher resolution by using electron microscopy, Professor Saibil’s team succeeded in estimating, for each perforin assembly, the number of molecules at each stage of the process. This confirmed a change from loosely packed small perforin assemblies on the membrane to larger and more tightly bound transmembrane pores. At the Peter MacCallum Cancer Centre in Melbourne, Ilia Voskoboinik highlights the medical importance of the results: “To kill virus-infected or cancerous cells, perforin must be quick and efficient, as achieved by the mechanism disclosed here. Patients born without functional perforin may present with fatal failure of the immune system and also have a higher risk of developing blood cancers.” His lab also performed experiments with lymphocytes that contained partially active perforin – variants such as those found in patients with the fatal human hyperinflammatory disease FHL. The results were consistent with the microscopic data obtained in London, and interestingly show that perforin pore formation overall is greatly hampered even if only a subpopulation of the perforin molecules are abnormal. With several stages of perforin pore formation now discovered, the next step will be to determine the corresponding molecular structures, which may then serve as targets for therapies that enhance or suppress the immune system as appropriate.
LCN/UCL/Physics/IPLS research in the national media with antimicrobial bullets from breast milk14/1/2016
|





 RSS Feed
RSS Feed