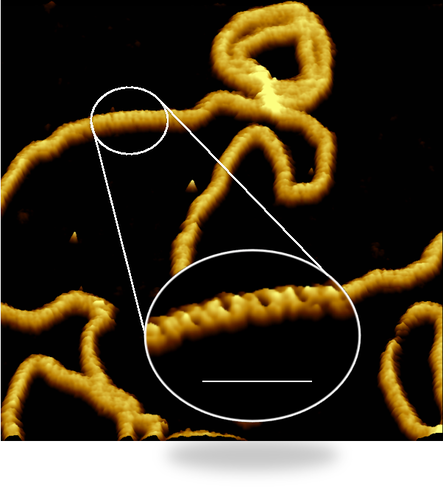
DNAThe Watson-Crick double-helix is the most iconic structure in biology, but DNA comes in a much richer variety of forms. Using atomic force microscopy, we can resolve the two strands of the double helix as they wind around each other in a single DNA molecule, and visualise proteins as they bind to DNA to, e.g., promote or repress the expression of genes. Together with Justin Molloy at the Francis Crick Institute and with AstraZeneca, we also aim to understand how cancer therapeutics affect DNA strand break repair mechanisms. Much of our DNA work has been done in close collaboration with our former PhD student and colleague Alice Pyne.
|
|
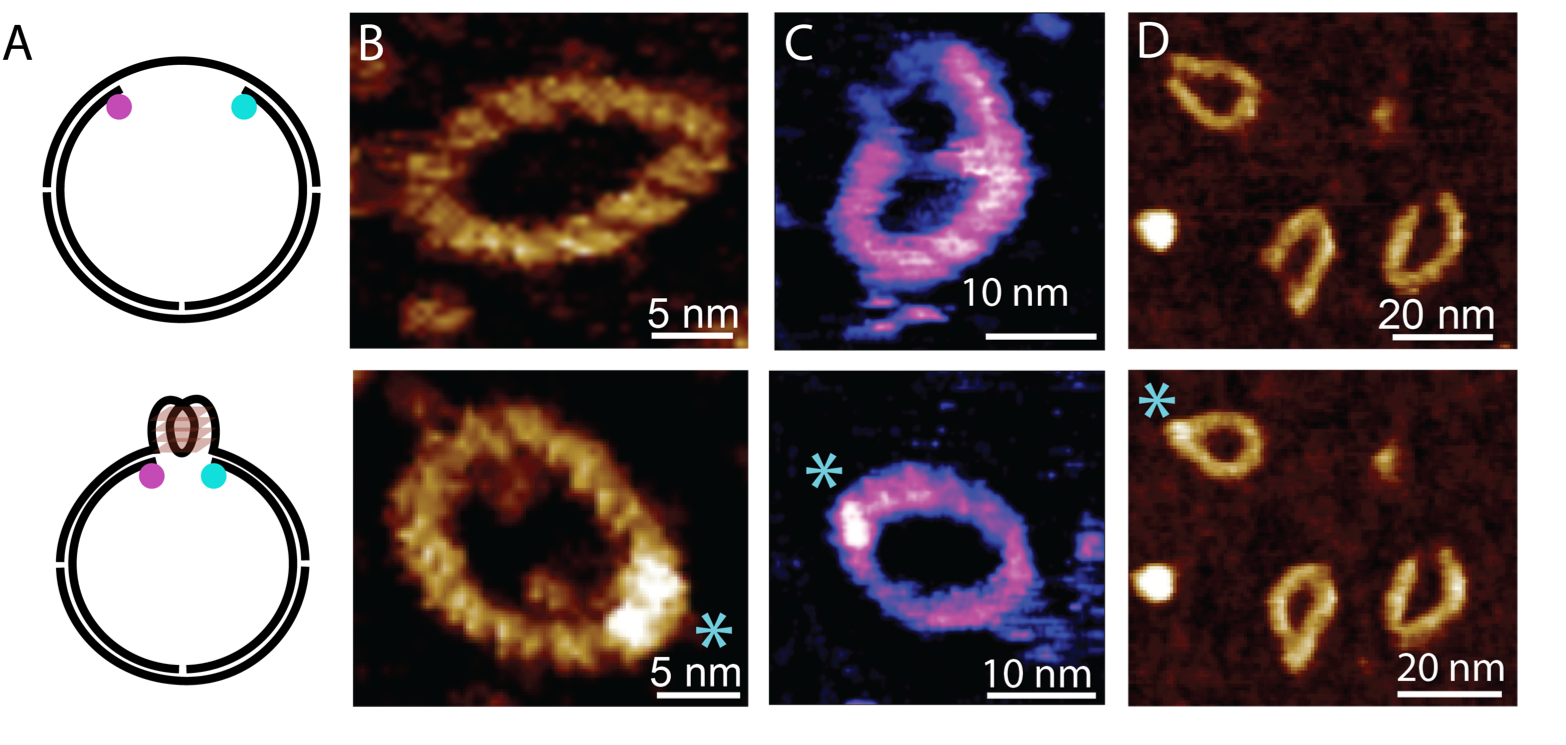
DNA is not always arranged in a double helix, also forming other structures such as structure fourfold DNA, which is found near DNA-encoded genes that are key in causing cancer. By the formation of such quadruplex DNA, the expression of cancer-related genes can be blocked. In collaboration with Ramon Vilar at Imperial College London, we have studied and visualised the formation of these quadruplexes in looped DNA that may be a better representation of DNA in the living cell than in the short and linear pieces of DNA that are more commonly used for such investigations.
|
Selected Publications
- N. A. W. Bell, P. J. Haynes, K. Brunner, T. M. de Oliveira, M. M. Flocco, B. W. Hoogenboom and J. E. Molloy, Science Advances 2021, 7, eabf3641
- A. L. B. Pyne, A. Noy, K. H. S. Main, V. Velasco-Berrelleza, M. M. Piperakis, L. A. Mitchenall, F. M. Cugliandolo, J. G. Beton, C. E. M. Stevenson, B. W. Hoogenboom, A. D. Bates, A. Maxwell and S. A. Harris, Nature Communications 2021, 12, 1053
- A. Pyne, R. Thompson, C. Leung, D. Roy, B. W. Hoogboom, Small, 2014, 10(16), 3257–3261